Array jobs: embarassingly parallel execution
Videos
Videos of this topic may be available from one of our kickstart course playlists: 2023, 2022 Summer, 2022 February, 2021 Summer, 2021 February.
Abstract
Arrays allow you to submit jobs and it runs many times with the same Slurm parameters.
Submit with the
--array=Slurm argument, give array indexes like--array=1-10,12-15.The
$SLURM_ARRAY_TASK_IDenvironment variable tells a job which array index it is.There are different templates to use below, which you can adapt to your task.
If you aren’t fully sure of how to scale up, contact us Research Software Engineers early.
More often than not, scientific problems involve running a single program again and again with different datasets or parameters.
When there is no dependency or communication among the individual program runs, these individual runs can be run in parallel on separate Slurm jobs. This kind of parallelism is called embarassingly parallel.
Slurm has a structure called job array, which enables users to easily submit and run several instances of the same Slurm script independently in the queue.
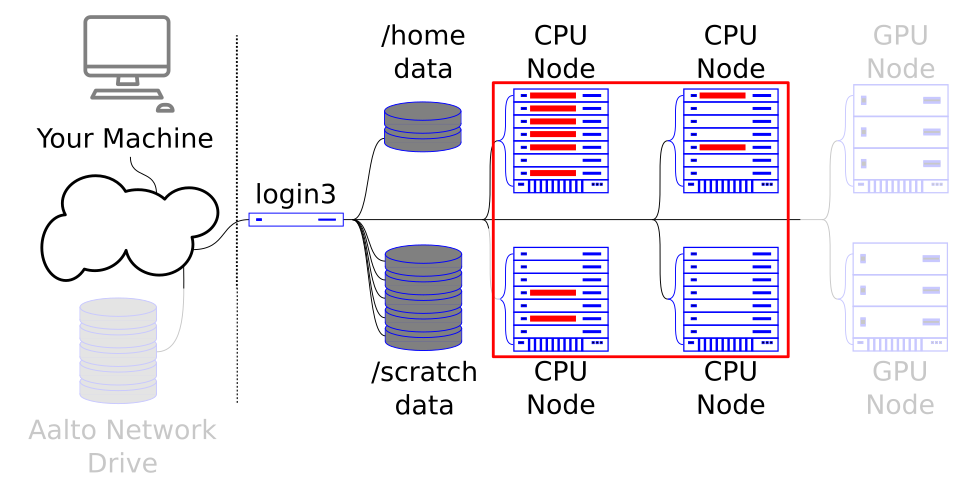
Array jobs let you control a large amount of the cluster. In Parallel computing: different methods explained, we will see another way.
Introduction
Array jobs allow you to parallelize your computations. They are used when you need to run the same job many times with only slight changes among the jobs. For example, you need to run 1000 jobs each with a different seed value for the random number generator. Or perhaps you need to apply the same computation to a collection of data sets. These can be done by submitting a single array job.
A Slurm job array is a collection of jobs that are to be executed with identical
parameters. This means that there is one single batch script that is to be run
as many times as indicated by the --array directive, e.g.:
#SBATCH --array=0-4
creates an array of 5 jobs (tasks) with index values 0, 1, 2, 3, 4.
The array tasks are copies of the submitted batch script that are automatically submitted
to Slurm. Slurm provides a unique environment variable SLURM_ARRAY_TASK_ID to each
task which could be used for handling input/output files to each task.
--array via the command line
You can also pass the --array option as a command-line argument to
sbatch. This can be great for controlling things without
editing the script file.
Important
When running array job you’re basically running identical copies of a single job. Thus it is increasingly important to know how your code behaves with respect to the file system:
Does it use libraries/environment stored in the work directory?
How much input data does it need?
How much output data does the job create?
For example, running an array job with hundreds of workers that uses a Python environment stored in the work disk can inadvertently cause a lot of filesystem load as there will be hundreds of thousands of file calls.
If you’re unsure how your job will behave, ask us Research Software Engineers for help for help.
Your first array job
Let’s see a job array in action. Lets create a file called
array_example.sh
and write it as follows.
#!/bin/bash
#SBATCH --time=00:15:00
#SBATCH --mem=200M
#SBATCH --output=array_example_%A_%a.out
#SBATCH --array=0-15
# You may put the commands below:
# Job step
srun echo "I am array task number" $SLURM_ARRAY_TASK_ID
Submitting the job script to Slurm with sbatch array_example.sh, you will get the message:
Submitted batch job 60997836
The job id in the message is that of the primary array job. This is common for all of the jobs in the array. In addition, each individual job is given an array task id.
As now we’re submitting multiple jobs simultaneously, each job needs an
individual output file or the outputs will overwrite each other. By default,
Slurm will write the outputs to files named
slurm-${SLURM_ARRAY_JOB_ID}_${SLURM_ARRAY_TASK_ID}.out. This can be overwritten using the
--output=FILENAME-parameter, when you can use wildcard %A for the
job id and %a for the array task id.
Once the jobs are completed, the output files will be created in your work directory,
with the help %u to determine your user name:
$ ls
array_example_60997836_0.out array_example_60997836_12.out array_example_60997836_15.out array_example_60997836_3.out array_example_60997836_6.out array_example_60997836_9.out
array_example_60997836_10.out array_example_60997836_13.out array_example_60997836_1.out array_example_60997836_4.out array_example_60997836_7.out array_example.sh
array_example_60997836_11.out array_example_60997836_14.out array_example_60997836_2.out array_example_60997836_5.out array_example_60997836_8.out
You can cat one of the files to see the output of each task:
$ cat array_example_60997836_11.out
I am array task number 11
Important
The array indices do not need to be sequential. For example, if after
running an array job you find out that tasks 2 and 5 failed, you can
relaunch just those jobs with --array=2,5.
You can even simply pass the --array option as a command-line argument to
sbatch.
More examples
The following examples give you an idea on how to use job arrays for different
use cases and how to utilize the $SLURM_ARRAY_TASK_ID environment
variable. In general,
You need some map of numbers to configuration. This might be files on the filesystem, a hardcoded mapping in your code, or some configuration file.
You generally want the mapping to not get lost. Be careful about running some jobs, changing the mapping, and running more: you might end up with a mess!
Reading input files
In many cases, you would like to process several data files. That is, pass
different input files to your code to be processed. This can be achieved by
using $SLURM_ARRAY_TASK_ID environment variable.
In the example below, the array job gives the program different input files,
based on the value of the $SLURM_ARRAY_TASK_ID:
#!/bin/bash
#SBATCH --time=01:00:00
#SBATCH --mem=1G
#SBATCH --array=0-29
# Each array task runs the same program, but with a different input file.
srun ./my_application -input input_data_${SLURM_ARRAY_TASK_ID}
Hardcoding arguments in the batch script
One way to pass arguments to your code is by hardcoding them in the batch script you want to submit to Slurm.
Assume you would like to run the pi estimation code for 5 different seed values, each
for 2.5 million iterations. You could assign a seed value to each task in you job array
and save each output to a file. Having calculated all estimations, you could take the
average of all the pi values to arrive at a more accurate estimate. An example of such
a batch script
pi_array_hardcoded.sh
is as follows.
#!/bin/bash
#SBATCH --time=01:00:00
#SBATCH --mem=500M
#SBATCH --job-name=pi-array-hardcoded
#SBATCH --output=pi-array-hardcoded_%a.out
#SBATCH --array=0-4
case $SLURM_ARRAY_TASK_ID in
0) SEED=123 ;;
1) SEED=38 ;;
2) SEED=22 ;;
3) SEED=60 ;;
4) SEED=432 ;;
esac
srun python slurm/pi.py 2500000 --seed=$SEED > pi_$SEED.json
Save the script and submit it to Slurm:
$ sbatch pi_array_hardcoded.sh
Submitted batch job 60997871
Once finished, 5 Slurm output files and 5 application output files will be created in your current directory each containing the pi estimation; total number of iterations (sum of iteration per task); and total number of successes):
$ cat pi_22.json
{"successes": 1963163, "pi_estimate": 3.1410608, "iterations": 2500000}
Reading parameters from one file
Another way to pass arguments to your code via script is to save the arguments to a file and have your script read the arguments from it.
Drawing on the previous example, let’s assume you now want to run pi.py
with different iterations. You can create a file, say iterations.txt
and have all the values written to it, e.g.:
$ cat iterations.txt
100
1000
50000
1000000
You can modify the previous script to have it read the iterations.txt
one line at a time and pass it on to pi.py. Here, sed is used
to get each line. Alternatively you can use any other command-line
utility, e.g. awk. Do not worry if you don’t know how sed works
- Google search and man sed always help. Also note that the line numbers
start at 1, not 0.
The script
pi_array_parameter.sh
looks like this:
#!/bin/bash
#SBATCH --time=01:00:00
#SBATCH --mem=500M
#SBATCH --job-name=pi-array-parameter
#SBATCH --output=pi-array-parameter_%a.out
#SBATCH --array=1-4
n=$SLURM_ARRAY_TASK_ID
iteration=`sed -n "${n} p" iterations.txt` # Get n-th line (1-indexed) of the file
srun python slurm/pi.py ${iteration} > pi_iter_${iteration}.json
You can additionally do this procedure in a more complex way, e.g. read in multiple arguments from a csv file, etc.
(Advanced) Two-dimensional array scanning
What if you wanted an array job that scanned a 2D array of points?
Well, you can map 1D to 2D via the following pseudo-code: x =
TASK_ID // N (floor division) and y = TASK_ID % N (modulo
operation). Then map these numbers into your grid. This can be
done in bash, but at this point you’d want to start thinking about
passing the SLURM_ARRAY_TASK_ID variable into your code itself for
this processing.
(Advanced) Grouping runs together in bigger chunks
If you have lots of jobs that are short (a few minutes), using array jobs may induce too much overhead in scheduling and you will create huge number of output files. In these kinds of cases you might want to combine multiple program runs into a single array job.
Important
A good target time for the array jobs would be approximately 30 minutes, so please try to combine your tasks so that each job would at least take this long.
Easy workaround for this is to create a for-loop in your Slurm script. For example, if you want to run the pi script with 50 different seed values you could run them in chunks of 10 and run a total of 5 array jobs. This reduces the amount of array jobs we need by a factor of 10!
This method demands more knowledge of shell scripting, but the end result is a
fairly simple Slurm script
pi_array_parameter.sh
that does what we need.
#!/bin/bash
#SBATCH --time=01:00:00
#SBATCH --mem=500M
#SBATCH --job-name=pi-array-grouped
#SBATCH --output=pi-array-grouped_%a.out
#SBATCH --array=1-4
# Lets create a new folder for our output files
mkdir -p json_files
CHUNKSIZE=10
n=$SLURM_ARRAY_TASK_ID
indexes=`seq $((n*CHUNKSIZE)) $(((n + 1)*CHUNKSIZE - 1))`
for i in $indexes
do
srun python slurm/pi.py 1500000 --seed=$i > json_files/pi_$i.json
done
Exercises
The scripts you need for the following exercises can be found in our
hpc-examples, which
we discussed in Using the cluster from a shell.
You can clone the repository by running
git clone https://github.com/AaltoSciComp/hpc-examples.git. Doing this
creates you a local copy of the repository in your current working
directory. This repository will be used for most of the tutorial exercises.
Array-1: Array jobs and different random seeds
Create a job array that uses the slurm/pi.py to calculate a
combination of different iterations and seed values and save them
all to different files. Keep the standard output (#SBATCH
--output=FILE) separate from the standard error (#SBATCH
--error=FILE).
Array-2: Combine the outputs of the previous exercise.
You find an pi-aggregate.py program in hpc-examples. Run this
and give all the output files as arguments. It will combine all
the statistics and give a more accurate value of \(\pi\).
Array-3: Reflect on array jobs in your work
Think about your typical work. How could you split your stuff into trivial pieces that can be run with array jobs? When can you make individual jobs smaller, but run more of them as array jobs?
(Advanced) Array-4: Array jobs with advanced index selector
Make a job array which runs every other index, e.g. the array can be indexed as 1, 3, 5… (the sbatch manual page can be of help)
Solution
You can specify a step function with colon and a number after indices.
In this case it would be: --array=1-X:2
See also
If you aren’t fully sure of how to scale up, contact us Research Software Engineers early. We are great at making these types of workflows.
For more information, you can see the CSC guide on array jobs
Please check the quick reference when needed.
What’s next?
The next tutorial is about shared memory parallelism.